- Products
- Analysis software module
- JSI medical systems
Sequencing software module SEQPATIENTanalysislaboratory
Add to favorites
Compare this product
Characteristics
- Function
- analysis
- Applications
- for sequencing, laboratory
Description
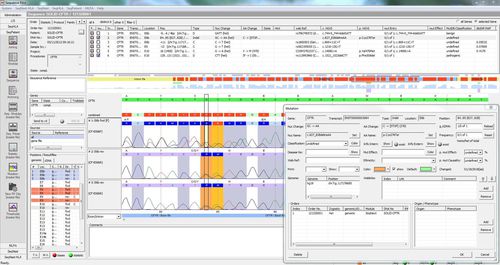
SEQPATIENT is a powerful and user-friendly application for alignment and variant detection of Sanger sequencing data. SEQPATIENT can analyse data from all common sequencing platforms. Visualisation of all detected variants like deletions, insertions, indels and SNPs is clear and intuitive. Analysis of genomic DNA and cDNA is possible. A peak area statistic function guarantees the detection of low frequency mutations like allelic dropouts, mosaics and somatic variants.
SEQPATIENT offers access to public SNP-databases like dbSNP, 1000 Genomes, COSMIC, ClinVar, ClinVitae, ExAC and gnomAD for classification and filtering. All result data can be exchanged with our variant database for Shared Experience And Knowledge - varSEAK, transferred to laboratory internal LIM Systems and / or issued as personalised patient reports.
Compatible with data from all common Sanger sequencing platforms
Easy setup of individual target regions via import of txt- or bed-files based on single downloaded genes (ENSEMBL / NCBI) or hg19 / hg38
Configurable base caller with sequencer dependent thresholds
Automatic and user defined trimming of sequencing result files
Patient identification and genotyping via SNP IDs
All results for one patient shown in one screen incl. found variants, amino acid changes, HGVS nomenclatures etc.
High sensitivity and specificity for detection of SNPs, deletions, insertions and indels
Internal mutation database keeping track of mutation detection history from SEQPATIENT and SEQNEXT
Catalogs
SEQUENCE PILOT
2 Pages
Other JSI medical systems products
Products
*Prices are pre-tax. They exclude delivery charges and customs duties and do not include additional charges for installation or activation options. Prices are indicative only and may vary by country, with changes to the cost of raw materials and exchange rates.










