- Laboratory
- Molecular biology
- Analysis software module
- JSI medical systems
HLA sequencing software module SEQNEXT-HLAanalysisfor mappinglaboratory
Add to favorites
Compare this product
fo_shop_gate_exact_title
Characteristics
- Function
- analysis, for mapping
- Applications
- laboratory, for HLA sequencing
Description
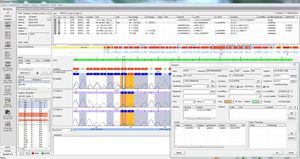
SEQNEXT-HLA is a powerful and user-friendly application for the analysis of HLA sequence based typing data from all common Next Generation Sequencing platforms and kits. All IMGT HLA database versions are available for download and import. For result calculation intron sequences can be regarded to exclude null alleles, moreover the phasing can be checked.
Warnings for coverages and background can be shown up. New alleles are called as well. The result can be displayed in maximum, 1 field or 2 field resolution and as NMPD code. All result data can be transferred to laboratory internal LIM Systems and / or issued as personalised patient reports.
Compatible with data from all common Next Generation Sequencing platforms and kits
Easy setup of individual target regions (ROIs) based on the IMGT HLA database
Analysis of fastq- or fastq.gz-files
Efficient standards for mapping, alignment, quality and allele calling
Configurable settings for allele calling
Analysis of complete loci (exon as well as intron data)
Detection / exclusion of null alleles based on intron sequences
Phasing check to remove ambiguities
Detection of new alleles
Catalogs
SEQUENCE PILOT
2 Pages
*Prices are pre-tax. They exclude delivery charges and customs duties and do not include additional charges for installation or activation options. Prices are indicative only and may vary by country, with changes to the cost of raw materials and exchange rates.